Biophysics
Insertion and Folding of Membrane Proteins
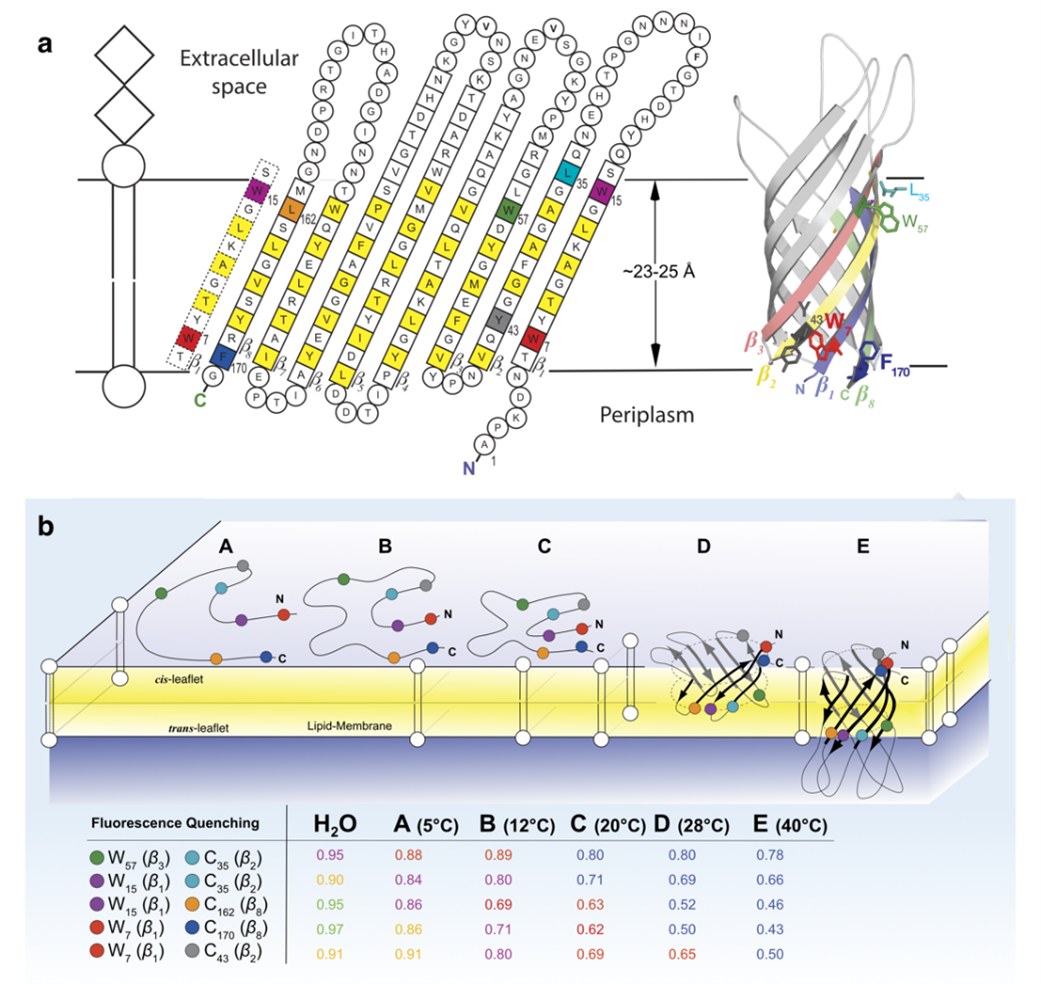
Biological membranes are essential structuring elements of all living cells. They are composed of a phospholipid bilayer that forms a hydrophobic barrier preventing the arbitrary exchange of solutes, and membrane-spanning as well as peripherally associated proteins. Membrane proteins allow the regulated exchange of solutes or the transduction of signals from one side of the membrane to the other or perform enzymatic or other functions. Although membranes are essential for all life, their biogenesis is not well understood and therefore this is the area of research for many cell biologists, biochemists, and biophysicists. There are only two types of transmembrane proteins that are distinguished by the secondary structure of their transmembrane domains, namely α-helical and β-barrel integral membrane proteins. The biophysical mechanisms of their insertion and folding into membranes are poorly understood. We have described major biophysical principles of the folding and membrane insertion of β-barrel membrane proteins from outer membranes (OM) of bacteria (OMPs) like Escherichia coli and also of the OM of mitochondria from cells of Homo sapiens [1]. These include structural characterizations of folding intermediates (Fig. 1B) of hVDAC1 and of the transmembrane domain of outer membrane protein A (OmpA, the amino acid sequence of OmpA is shown in Figure 1A) and interactions of bacterial OMPs (OmpA, OmpG, BamA, FomA, NalP etc.) with molecular chaperones from the bacterial periplasm, like Skp, SurA, and FkpA. For these investigations, we have developed a range of novel methods and approaches, see e.g. [2, 3]
While folding and membrane insertion of β-barrel proteins into pure lipid bilayers, detergent micelles or even amphipathic polymers can be spontaneous, e. g. from a denatured form of the β-barrel membrane protein in solutions of chaotropic reagents like urea, a heteropentameric protein complex from the OM of E. coli has been identified in recent years, named barrel assembly machinery (BAM) complex that that is required for insertion and folding into cellular OMs. The OM of E. coli has a lipid composition that prevents spontaneous folding and insertion. Therefore, the BAM complex is essential for cell viability. The physical principles by which the BAM-complex works, are currently a major focus of research of the Biophysics group. We have isolated all five proteins of the BAM-complex of E. coli and examined their function in membrane protein folding of our main model protein for folding studies, OmpA. Only two subunits of the BAM complex have been reported as essential for survival of E. coli under laboratory conditions, BamA (~800 amino acid residues) and BamD (~240 residues). However, E. coli also does not tolerate the simultaneous deletion of BamB and BamE. On the other hand, certain bacteria like Thermus thermophilus only contain BamA, which is evolutionarily conserved, but not the other 4 subunits found in E. coli. These observations suggest that BamA is central to the function of the BAM complex from E. coli and that the other 4 subunits serve specialized functions. Through biophysical analyses, we recently demonstrated several key features of the BAM complex from E. coli. Among these are:
- The three main components of the BAM-complex, BamA, BamB, and BamD are all able to individually accelerate the kinetics of folding of OmpA into model membranes, hinting that these subunits are involved in sequential rather than parallel steps that need concerted action.
- We could devise a membrane model system that allows us to trap folding intermediates of OmpA (and likely of other OMPs) bound to BamA. This enables the identification of their local structures and also of their local contacts with BamA.
- We have identified and characterized direct interactions of the molecular chaperone Skp from the periplasm of E. coli with BamD and BamA.
- We established the binding regions of BamB to membrane lipids, demonstrating that BamB is not only anchored to the membrane by its N-terminus, but also via a separate lipid binding domain.
These are key results to determine the roles and the functions of the entire BAM complex of E. coli as well as of its individual subunits. They are also a solid basis to examine BAM-interactions with several OMPs of importance in Gram-negative bacteria like E. coli. This is not only of interest to basic research, but also for biotechnological applications in view of a growing number of multi-drug resistant strains of pathogenic Gram-negative bacteria. The basic research may provide a basis for new classes of antibiotics that render the BAM-complex or important client OMPs dysfunctional, killing the bacteria.
[1] J.H. Kleinschmidt, Biochim Biophys Acta 1848 (2015) 1927-1943.
[2] A. Schüßler, S. Herwig, J.H. Kleinschmidt, Methods Mol Biol 2003 (2019) 145-162.
[3] L. Gerlach, O. Gholami, N. Schürmann, J.H. Kleinschmidt, Methods Mol Biol 2003 (2019) 465-492.
Prof. Dr. Jörg H. Kleinschmidt
full member

- Telephone
- +49 561 804-4041
- jhk[at]uni-kassel[dot]de
- Location
- Universität Kassel
Fachbereich 10 - Naturwissenschaften & Mathematik
Institut für Biologie
Heinrich-Plett-Straße 40
34132 Kassel
- Room
- 2249
